In the field of molecular biology, the ability to accurately quantify nucleic acids is essential for numerous applications, ranging from basic research to clinical diagnosis. While traditional quantitative PCR (qPCR) has been the most useful technique for nucleic acid quantification, recent advances have led to the emergence of an innovative technology: digital droplet polymerase chain reaction (ddPCR). Offering unparalleled precision, sensitivity, and absolute quantification capabilities, ddPCR has revolutionized the landscape of nucleic acid analysis.
Droplet digital PCR (ddPCR) represents a groundbreaking advancement in nucleic acid quantification, offering unparalleled precision, sensitivity, and absolute quantification capabilities. At the heart of ddPCR is sophisticated instrumentation designed to split samples into discrete droplets, allowing precise quantification of target DNA or RNA molecules.
What is ddPCR?
Droplet digital PCR (ddPCR) represents the pinnacle of innovation in nucleic acid quantification. In essence, ddPCR is an advanced iteration of Traditional PCR, employing the novel strategy of dividing a sample into thousands of tiny droplets, each acting as an independent reaction chamber. This partitioning allows absolute quantification of target DNA or RNA molecules, which is unachievable by conventional PCR methods.
Steps of ddPCR
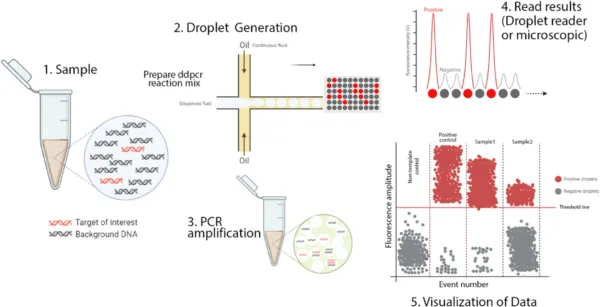
The ddPCR process works through the following steps:
Sample partition:
The sample is divided using a water-oil emulsion or microfluidics, generating numerous individual reactions within different droplets.
Amplification:
Within each droplet, PCR amplification occurs, facilitated by the inclusion of PCR reagents such as primers, nucleotides, and DNA polymerase.
Endpoint Analysis:
After amplification, the droplets are examined by fluorescence detection methods to determine the presence or absence of amplified target sequences.
Digital Quantization:
Fundamentally, dd PCR achieves digital quantification by counting the number of positive and negative droplets, allowing accurate determination of the concentrations of target molecules in the original sample.
Data analysis:
The acquired fluorescence data are meticulously analyzed to calculate absolute concentrations of target nucleic acid, a hallmark of ddPCR precision and reliability.
Discovery and evolution of ddPCR
The genesis of droplet PCR dates back to the pioneering work of researchers at Raindance Technologies and the National Institute of Standards and Technology (NIST), USA. In the early 2000s, Dr. Fred Kramer and his team They embarked on a journey to conceptualize and refine the principles of split PCR, laying the foundation for the eventual realization of ddPCR.
A pivotal moment in the trajectory of ddPCR came with the publication of Seminal article by Hindson et al. in Analytical Chemistry in 2011. This landmark study elucidated the microfluidic emulsification technology critical for splitting samples into droplets, marking a watershed moment in the evolution of ddPCR. Raindance Technologies' subsequent efforts propelled ddPCR into the scientific spotlight, facilitating its widespread adoption in research laboratories around the world.
Principles of ddPCR
He beginning The foundation of ddPCR encapsulates a fusion of traditional PCR methodologies with innovative digital quantification capabilities. The meticulous partitioning of samples into discrete droplets serves as the cornerstone of ddPCR's precision and accuracy. By enabling absolute quantification without relying on standard curves, ddPCR transcends the limitations of traditional qPCR and offers researchers unparalleled insights into nucleic acid dynamics.
The advantages that ddPCR confers over traditional qPCR methodologies are multiple:
- Improved Accuracy: ddPCR heralds a new era of precision in nucleic acid quantification, particularly at low concentrations, mitigating variability and improving precision.
- Improved sensitivity: The increased sensitivity of ddPCR allows researchers to detect and quantify minute amounts of target molecules, opening perspectives for the detection of rare mutations and the analysis of viral nucleic acids.
- Resistance to PCR inhibitors: The resistance of ddPCR to PCR inhibitors underscores its utility in the analysis of complex or impure samples, ensuring reliable quantification even in challenging experimental contexts.
- Absolute Quantification: Unlike qPCR's reliance on relative quantification, droplet digital PCR facilitates absolute quantification, eliminating the need for external calibration and improving the robustness of results.
- Reduced variability: By dividing samples into individual droplets, ddPCR mitigates variability resulting from pipetting errors and sample-to-sample variations, generating reproducible and reliable data.
- Multiplexing Capability: The multiplexing prowess of ddPCR allows researchers to simultaneously quantify multiple targets within a single sample, optimizing assay workflows and conserving resources.
- Applications in Liquid Biopsy and Clinical Diagnosis: The unparalleled sensitivity of ddPCR makes it indispensable in liquid biopsies and clinical diagnostics, facilitating the detection of circulating tumor DNA and genetic mutations with unprecedented precision.
Limitations of ddPCR
However, despite its transformative potential, ddPCR is not without limitations:
- Cost: The initial capital investment and per-sample costs associated with ddPCR instrumentation and consumables may pose barriers to adoption, particularly for laboratories with limited resources.
- Performance: ddPCR systems typically exhibit lower throughput compared to traditional qPCR platforms, limiting their scalability for high-throughput applications.
- Preparation of complex samples: The complexities of droplet generation and sample partitioning in ddPCR require meticulous attention to experimental protocols, potentially impeding workflow efficiency.
- Limited dynamic range: The dynamic range of ddPCR may be limited, requiring dilution or concentration steps for accurate quantification of target molecules that fall outside the linear range of detection.
- Challenges of trial design: Designing optimized assays requires careful consideration of factors such as droplet stability and PCR conditions, demanding experience, and iterative optimization.
- Complexity of data interpretation: Data interpretation requires sophisticated analytical tools and computational expertise, particularly for nuanced data sets or complex experimental designs.
- Multiplexing restrictions: Although ddPCR allows for multiplexing, multiplexing capacity may be restricted compared to qPCR, requiring judicious assay design and optimization.
Conclusion
In conclusion, droplet digital PCR is a model of innovation in nucleic acid quantification, offering unprecedented precision, sensitivity, and absolute quantification capabilities. From its inception to widespread adoption across diverse research domains, it exemplifies the power of interdisciplinary collaboration and technological ingenuity to drive scientific progress. While its limitations are recognized, the transformative potential of ddPCR to elucidate the complexities of nucleic acid dynamics remains unequivocal, paving the way for a new era of precision molecular biology and clinical diagnosis.













Leave feedback about this